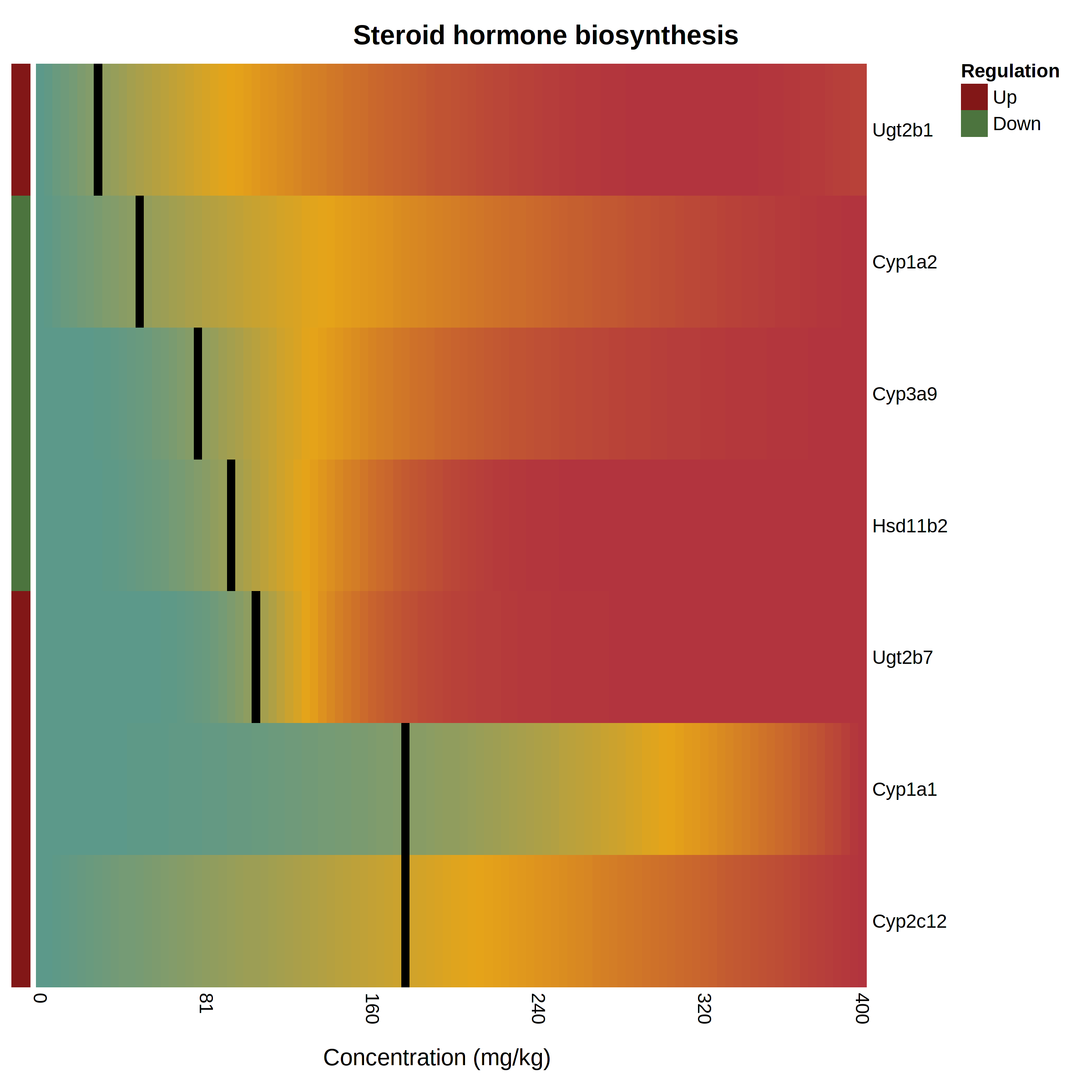
The pathway heatmap is an appealing visualization to clearly shows how the expression of each pathway gene compares to the others. It is generated when you click a pathway or gene set name in the “Gene Set Enrichment” panel at the result page. An example output is shown below The pathway heatmap values are calculated through a series of steps: The fitted model for each gene is evaluated across the range of doses in the uploaded data. The resulting modeled expression values are normalized
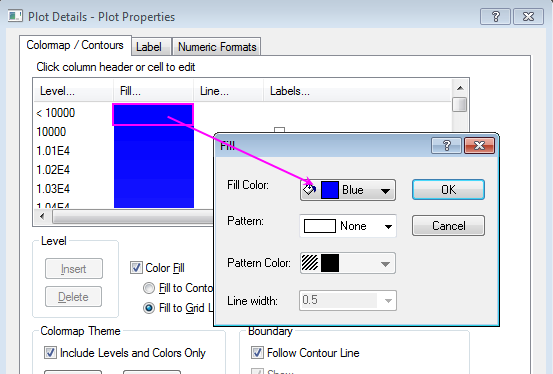
Help Online - Tutorials - Create Heatmap with Virtual Matrix and

Heatmap of the fold change of SCMs between YF-6 and GF-6. The

Heat map of cluster analysis of mouse serum metabolites and
Mouseflow - How to Interpret a Heatmap

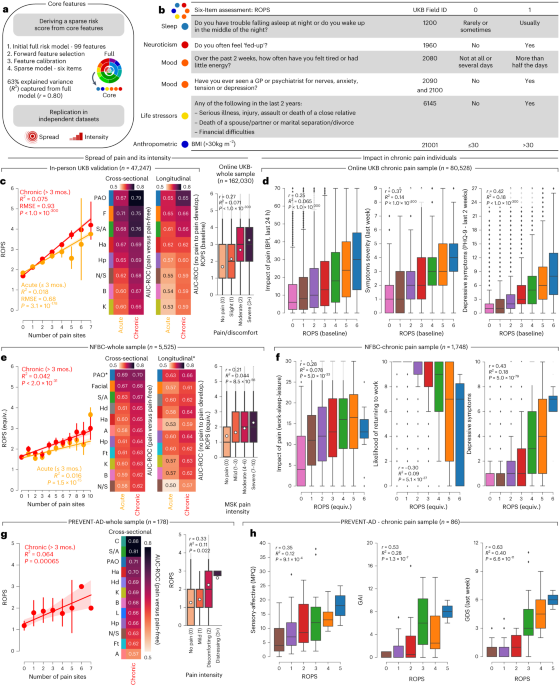
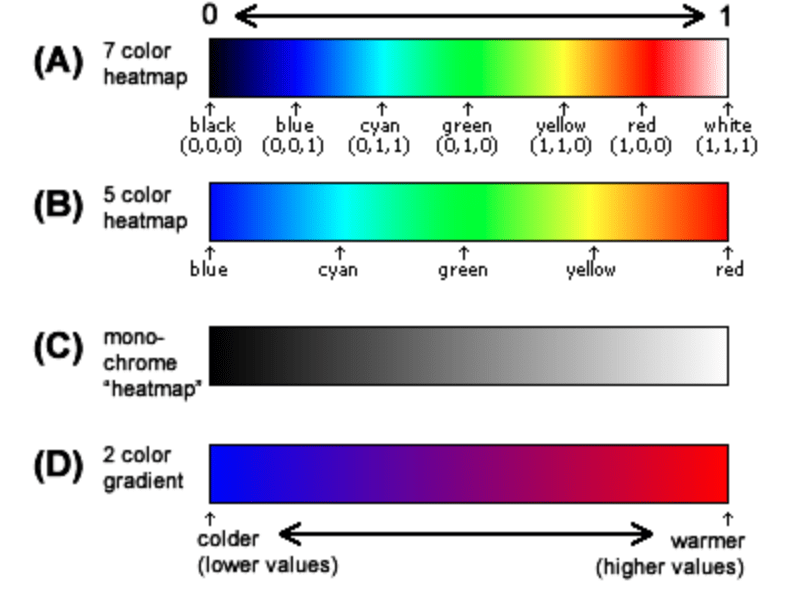
Dose-response metabolomics and pathway sensitivity to map
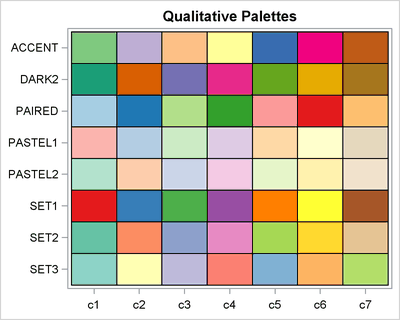
How to choose colors for maps and heat maps - The DO Loop

Dose-response metabolomics and pathway sensitivity to map
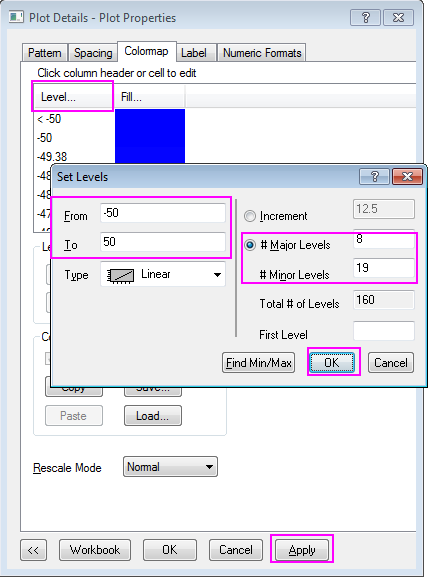
Help Online - Tutorials - Create Heatmap with Virtual Matrix and

What do the colours on the pathway heatmaps represent? - FastBMD - OmicsForum
Choose a good heatmap color scale with viridis (Revolutions)
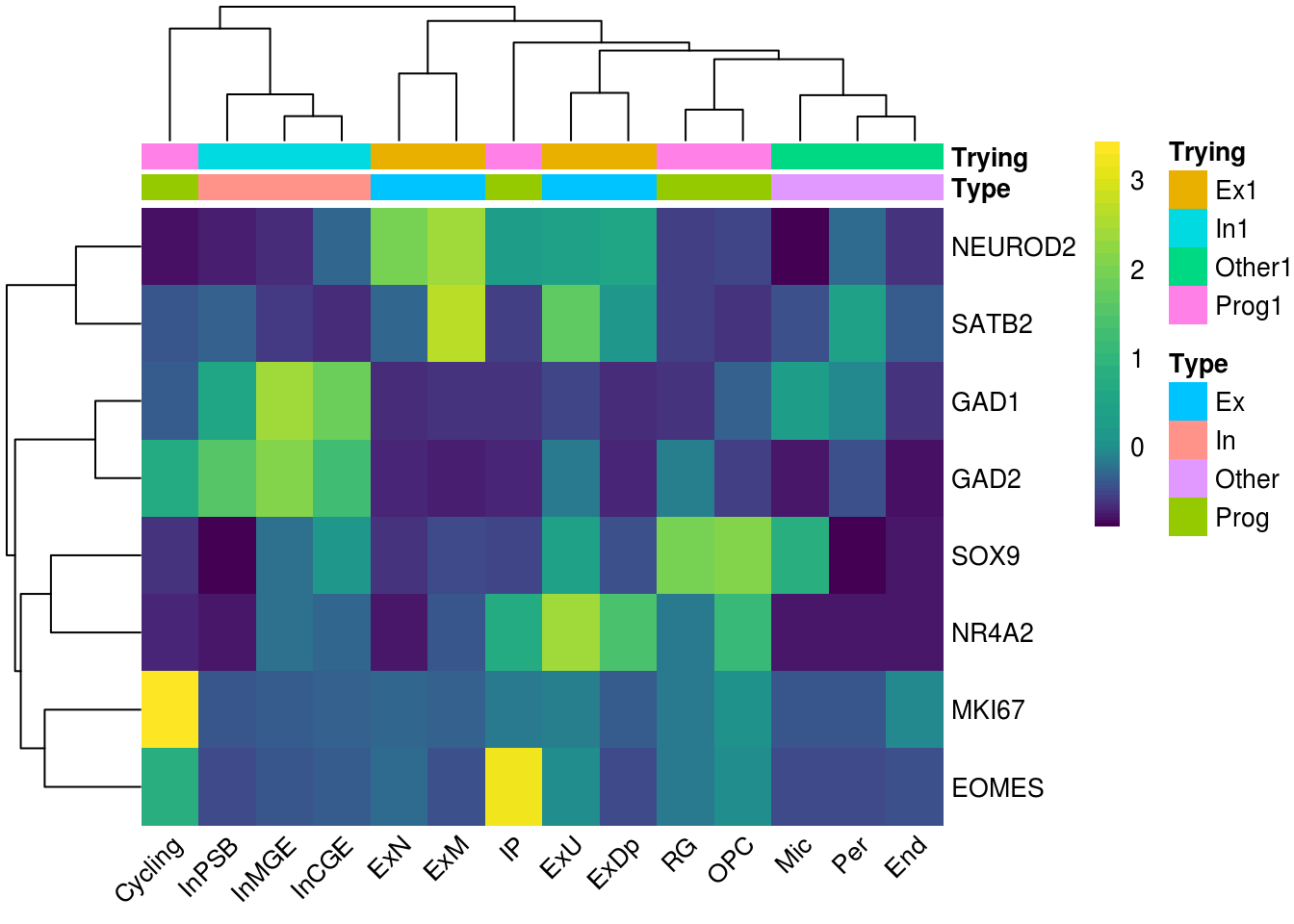
Chapter 9 Heatmap Color Palette Single Cell Multi-Omics Data

A Heatmap of-log10-transformed P-values for all 32 significant CP