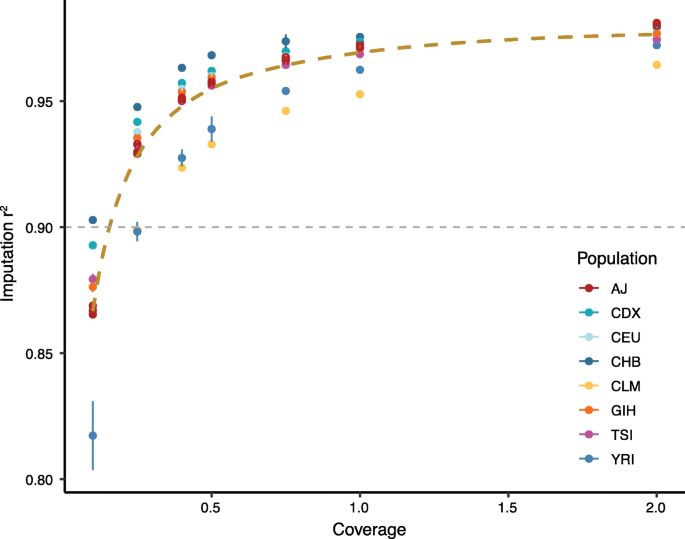
EM-PCA for Ultra-low Coverage Sequencing Data. Contribute to Rosemeis/emu development by creating an account on GitHub.

PDF) Large-scale Inference of Population Structure in Presence of Missingness using PCA

Emu: species-level microbial community profiling of full-length 16S rRNA Oxford Nanopore sequencing data

PDF) Detecting Selection in Low-Coverage High-Throughput Sequencing Data using Principal Component Analysis

GitHub - kasumaz/AdultOLgenesis: Single cell analysis of the generation and regulation of oligodendrocyte lineage cells from the adult subventricular zone.
GitHub - FranckLejzerowicz/microbiome_analyzer: Command line tool that writes commands to run one by one to perform analyses (mainly using qiime2) on a HPC running Slurm, Torque or none of these.

Deep representation features from DreamDIAXMBD improve the analysis of data-independent acquisition proteomics
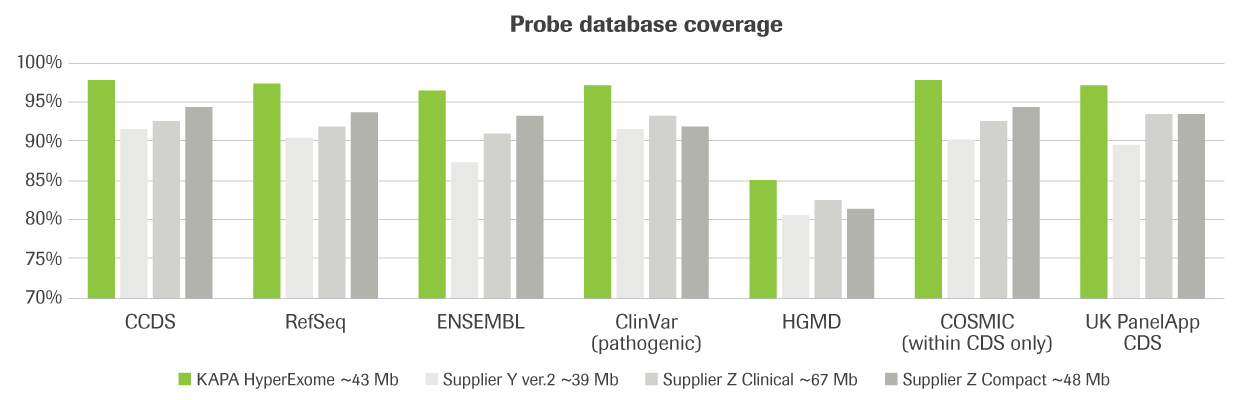
KAPA HyperExome - Roche Sequencing Store
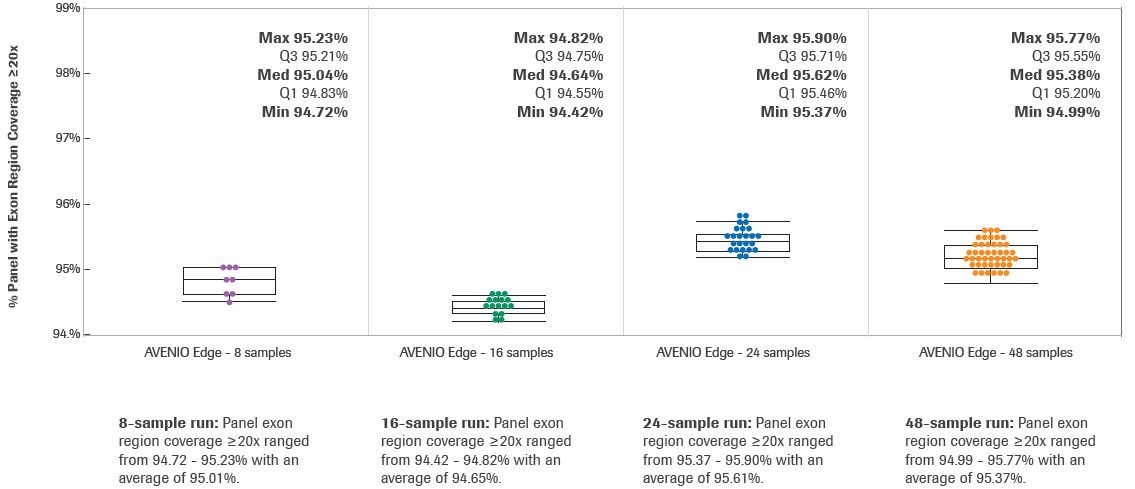
Using HyperExome probes for DNA hybridization workflows
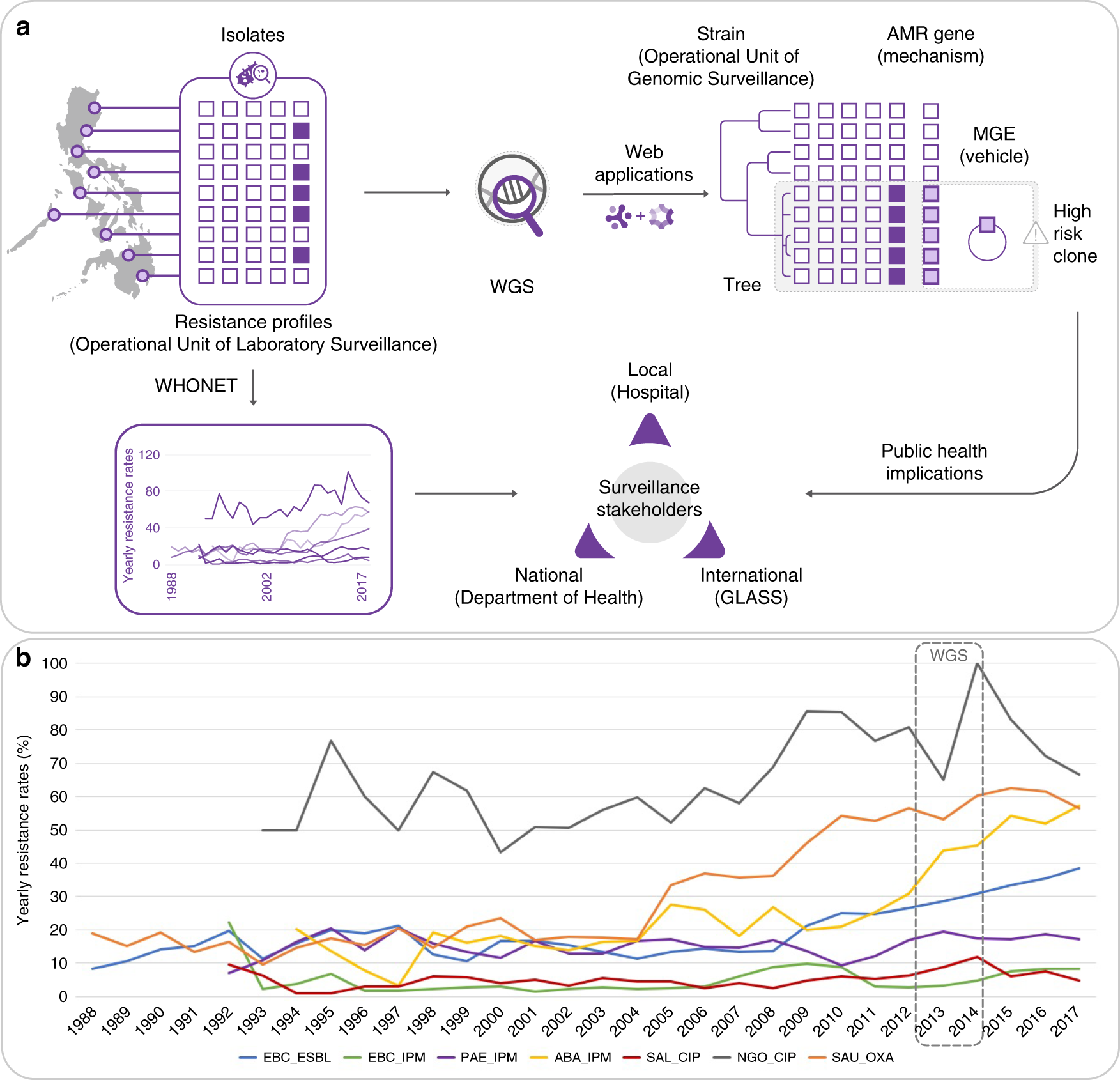
Integrating whole-genome sequencing within the National Antimicrobial Resistance Surveillance Program in the Philippines

Expressed Exome Capture Sequencing (EecSeq): a method for cost-effective exome sequencing for all organisms with or without genomic resources

Systematic dissection of biases in whole-exome and whole-genome sequencing reveals major determinants of coding sequence coverage

A hierarchical clustering of individuals from the HapMap, HGDP and TGP
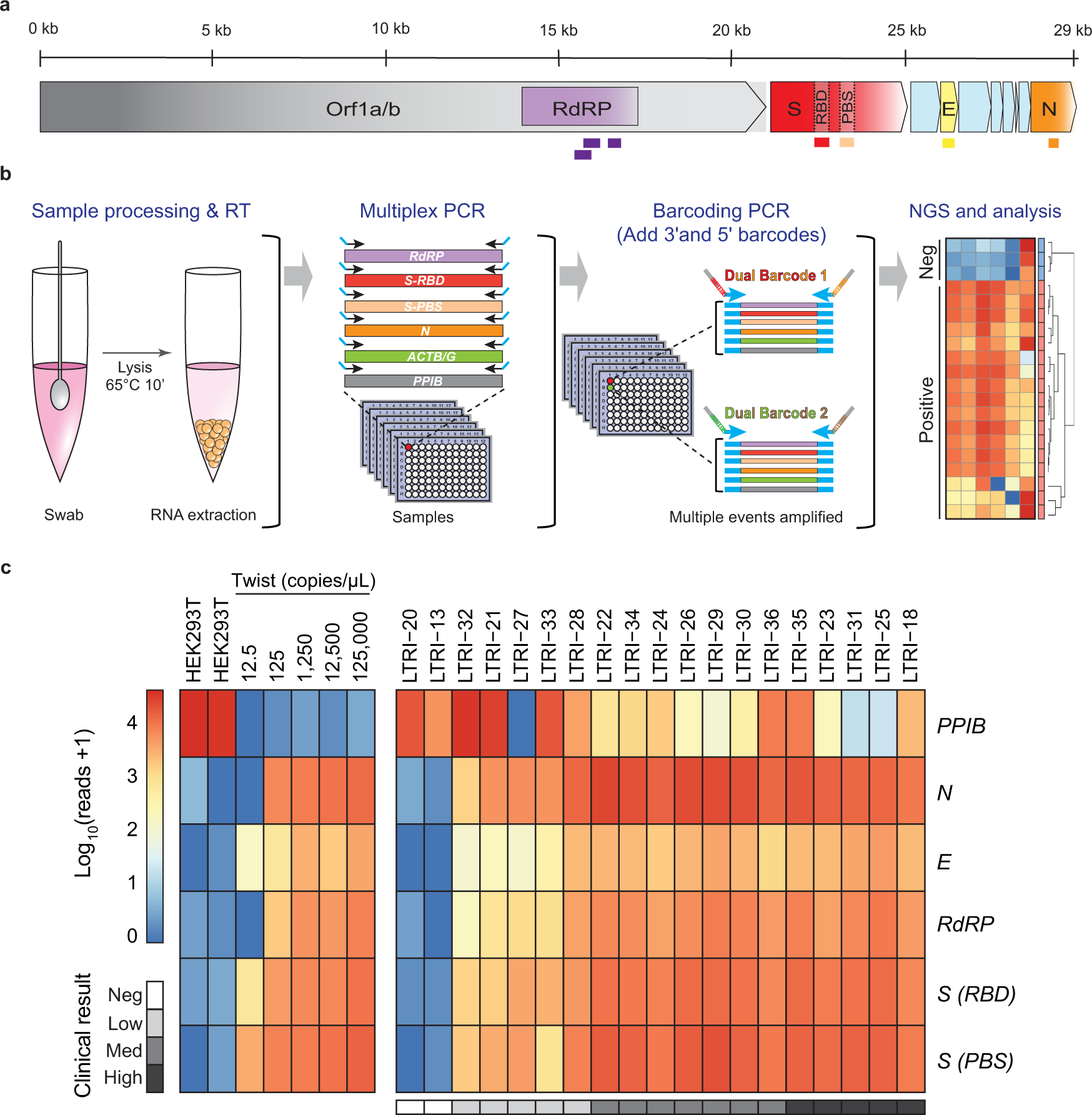
A multiplexed, next generation sequencing platform for high-throughput detection of SARS-CoV-2

PDF) Large-scale Inference of Population Structure in Presence of Missingness using PCA